Completing the puzzle of life on Earth
Behind the scenes of Sanger's Tree of Life programme
In 1998, a millimetre-long nematode worm, Caenorhabditis elegans, was the first animal to have its whole genome sequenced. This achievement laid the groundwork for the first human genome five years later. Now an effort is underway to revolutionise biology itself – by sequencing not just one or two species, but all of them
Why sequence all life on earth?
The Sanger Institute was founded to complete the genome of C. elegans and Homo sapiens. Now it is at the forefront of a new endeavour, forging a path where none exists. Together with ten partners, the Institute has embarked on a vast new voyage of discovery: the Darwin Tree of Life project, which aims to sequence the genomes of around 70,000 animal, plant, fungi and protist species in Britain and Ireland. These genomes will in turn contribute to the Earth BioGenome Project (EBP), the global effort to sequence the genomes of all species on our planet.
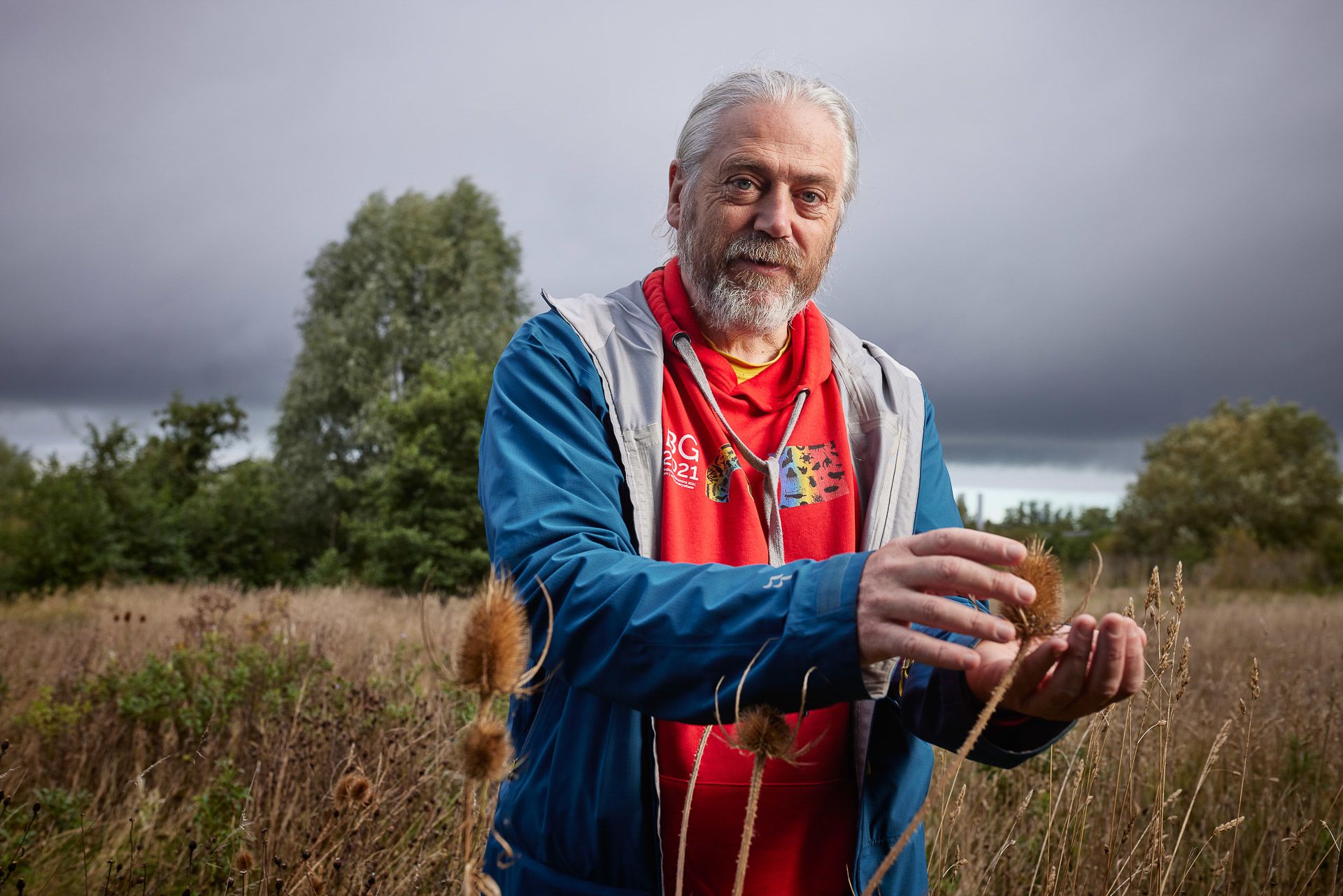
For Mark Blaxter, Head of the Tree of Life Programme at Sanger, this is his dream job. “When the Human Genome Project was completed, I wrote an article that said, ‘Great, the human genome is done… now we can do the rest of life.’ Ever since I’ve been a scientist, I’ve wanted to observe life at the genomic level. Much of our understanding of life is based on a handful of organisms – sequencing all life will enable us to properly understand evolution and will be the foundation of biology for the next century.”
Darwin Tree of Life is similar to the Human Genome Project in that the precise applications and discoveries it will enable are not yet fully known – it is a moonshot endeavour that requires technological and experimental advances along the way if it is even to be completed. And in a world facing the twin existential threats of biodiversity loss and global heating, it is perhaps natural to ask what sequencing the genomes of all the planet’s species will do to address these problems.
“DNA is the basis of all complex life on earth, the genetic machinery is the same whether we are talking about a human being or a sponge or a fern. So understanding how DNA is arranged into genomes, and how genomes have evolved over millions of years, is really the key to understanding everything.”
Mark Blaxter

Though this genetic machinery is common to all life, the sheer range of forms and abilities it can create is astounding. We need only to look at the natural world to see this, whether that is the cancer-proof, age-defying biology of a naked mole-rat, the reactive camouflage of a cuttlefish or the regenerative abilities of plants. All of these abilities are the result of genes and the proteins they create.
There are also large regions of DNA that do not code for proteins, which account for around 99 per cent of the genome in humans. Some of these regions are involved in gene regulation, but most of this 'dark matter' of the genome is likely neutral, invisible to the forces of evolution. About half of our genome is made up of remnant mobile DNA elements that generate additional copies of themselves but individually do little harm.
But more recently, research has begun to explore the possibility that dark matter DNA can be recruited to new functions, such as the ‘antifreeze’ gene that prevents the blood and tissues of Arctic cod from freezing in conditions that would kill other fish, which researchers now think evolved independently from a dark region of the genome, rather than from an existing gene.
It is this untapped promise that is at the heart of Darwin Tree of Life and the reason why reference genomes – complete from end to end of every chromosome – are so crucial. These complete genomes hold the key not only to what makes a species what it is, but how it got there and what it could become in future.
Digging beneath the surface of ecology
All life is arranged into ecosystems and, just like any other species, humans depend on them for survival. Understanding how ecosystems work – and how they can break down – will be one of the benefits of extensive genomic data.
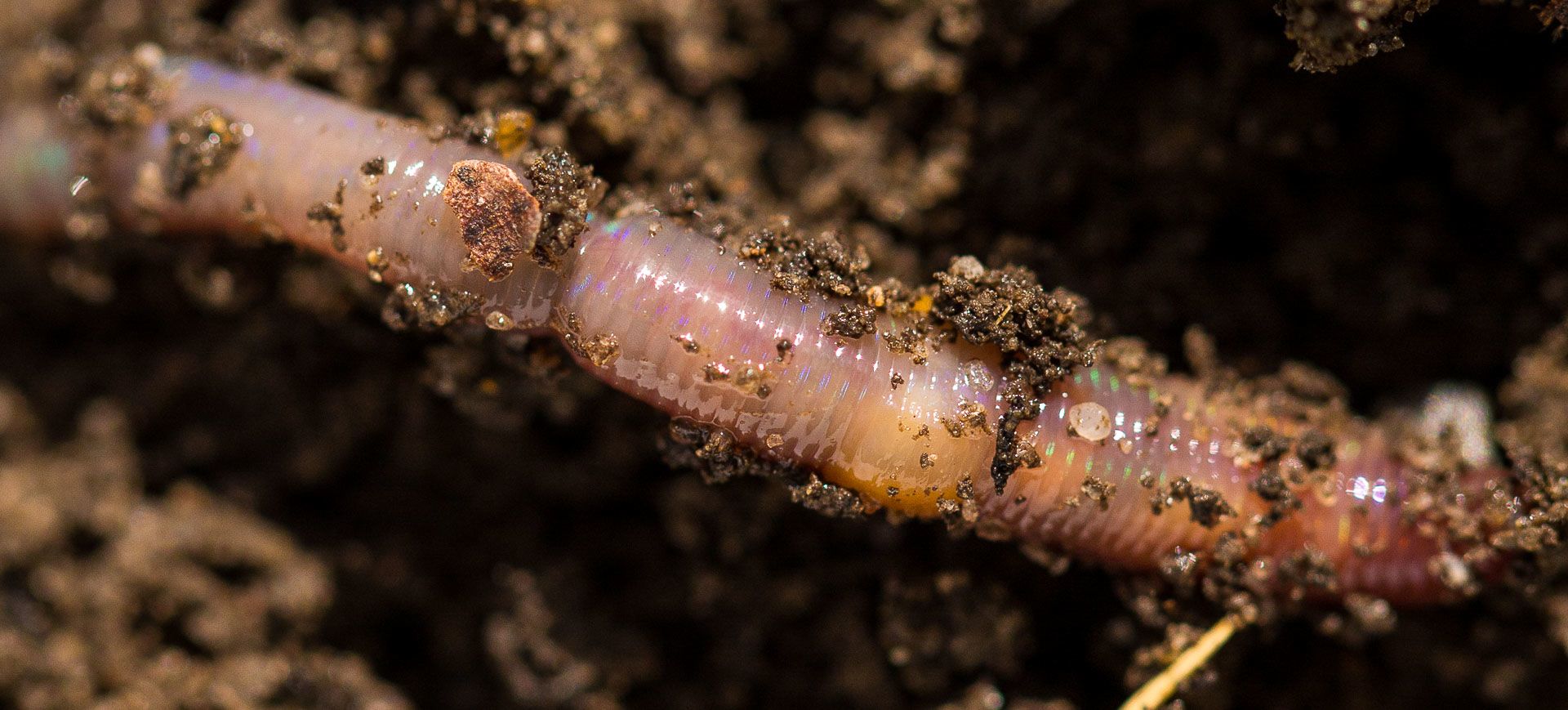
Take soil, for example. Most of the food the world produces comes from soil. It is home to around a quarter of land species. It is a vast repository of carbon and water. At least, it should be – but in many places these functions have been steadily eroded by the way humans use the land. A recent study in England found that 42 per cent of fields were deficient in earthworms, with some fields having none at all. Earthworms renew the soil and without them it loses many of its capabilities, including to produce crops. What we don’t know is what’s behind the decline.
Farming practices are certainly a factor, but what about the role of pesticides like neonicotinoids? One species of earthworm, Eisenia fetida, has historically been used to evaluate the toxicity of agricultural chemicals. If it was deemed ‘safe’ for E. fetida, it was deemed safe for all species. But even among the 27 native British earthworm species, sensitivity to chemicals varies. And what about the billions of other microorganisms that live in every spoonful of soil, the fungi and bacteria that also contribute to soil health?
“These genomes are going to have some really major positive impacts on the relationship between humans and the rest of life on the planet,” said Mark. “We should be able to start conserving whole ecosystems, rather than just a single target species – to understand how all the species interact, how to put an ecosystem that has been degraded back together again.”

Conserving endangered species
While some of the benefits of genomic data will take years to be realised, some genomes are already having a direct impact on conservation and biodiversity.
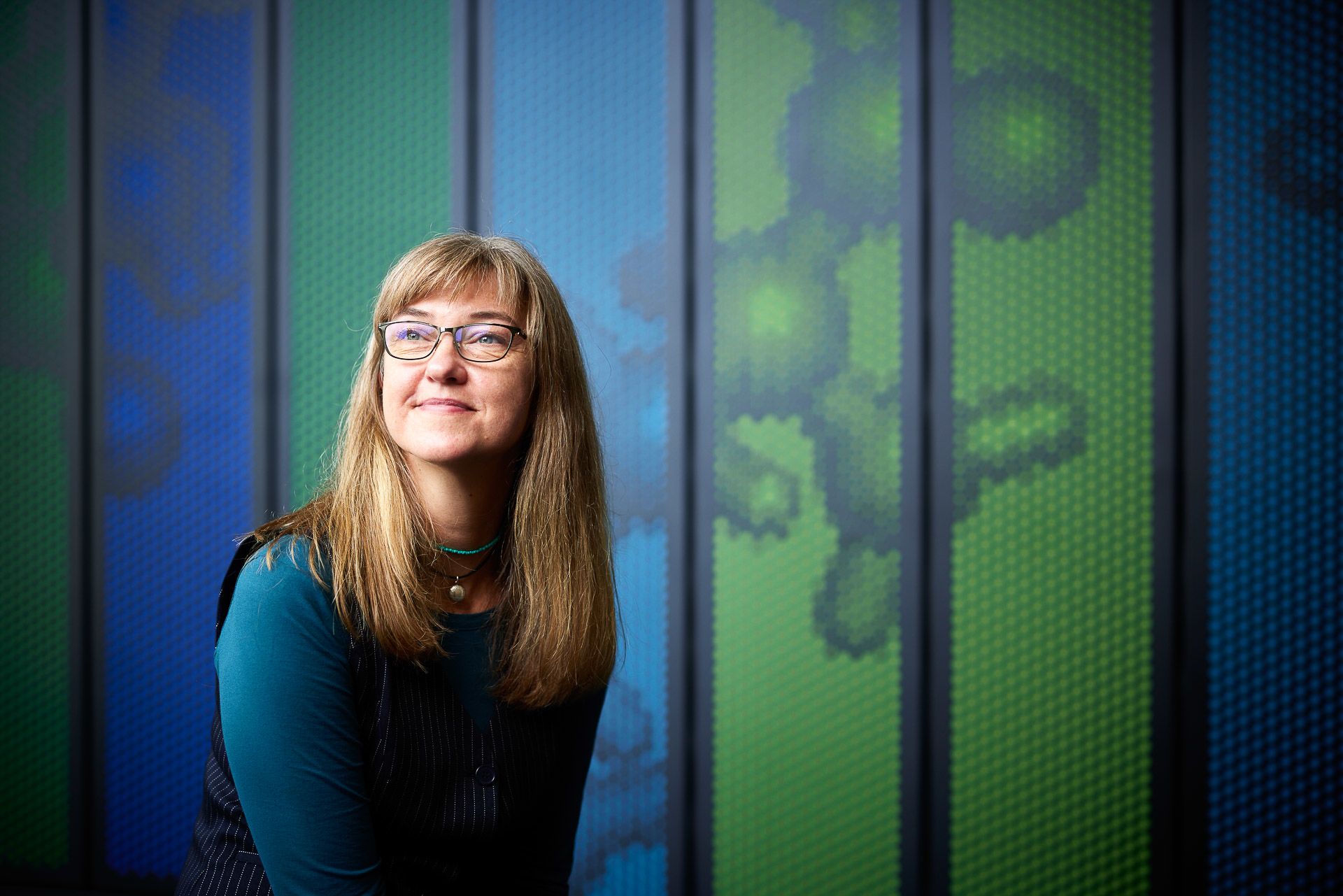
Kerstin Howe is Head of Production Genomics on the programme. Her career goes back to the Human Genome Project and since then she has helped to create genomes that are being used to protect some of the world’s rarest species. “I’m involved in the Vertebrate Genomes Project, which has helped to pave the way for Darwin Tree of Life. One of the first genomes we created was for the kākāpō, a parrot endemic to New Zealand. There are just over two hundred kākāpō left, up from 51 in 1995. While this is a conservation success, we need to make sure we preserve as much genetic diversity in the species as we can to avoid inbreeding and give them the greatest chance of recovery. The kākāpō genome allows us to do that.”
In Britain, the same application will be used to help preserve fragmented populations of the native red squirrel, one of the first genomes in the project to be completed, and the water vole, which is endangered in the country.
Perhaps genomics can also help us to understand why the twite, a finch that was abundant in English uplands just 20 years ago, is on the verge of extinction in England despite sustained conservation efforts. Excitingly, the data open the door to genomically-informed ecosystem conservation, where the genomes of all the species in an ecosystem can be examined to understand how they mesh together and contribute to ecosystem resilience and recovery.

Fuelling economies of the future
It is envisaged that whole new economies will be founded on genomic data. Among them are new biomaterials, pharmaceuticals, fuels and crops, and some of these are closer to reality than you might think.
Plastic is a huge problem from an environmental and health perspective. An island of plastic waste in the Pacific Ocean covers an area three times the size of France. Because plastic breaks down into smaller pieces rather than biodegrading, microplastic particles have polluted every ecosystem on earth, including the placentas of unborn babies, with the long-term effects unknown.
Ironically, the other problem with plastic is that we will run out of the raw ingredients required to make it in the coming years as the world phases out oil.
“Researchers are already figuring out how to replace oil-derived plastics with materials created from or inspired by spider silk,” said Mark. “What gives a particular silk its remarkable strength, or elasticity? The answer is in the spider’s genome and if we can understand that biology, we can recreate it using engineered bacteria in bioreactors, fed by renewable resources. This is a great example of how genomes can become the raw material of whole new bio-economies.”
Another example from the University of Edinburgh shows how genomics is being used to tackle greenhouse gas emissions at source. By analysing the genotype of individuals, the programme is selectively breeding ‘green cows’ that require less feed and produce less methane.
What does it take to create a reference genome?

To put the scale and ambition of the Darwin Tree of Life project in perspective, it’s worth remembering that it took over a decade to create the human reference genome. In the two decades since, scientific and technological advances have massively increased the speed at which DNA can be sequenced. But 70,000 is a big number, and to date just over a thousand species have high-quality reference genomes.
It is also important to note that sequencing a genome for the first time is very different to sequencing the genomes of many individuals from the same species. One might assume that the human genome, the blueprint for the most advanced species on earth, would be the most complex. This assumption would be wrong.
In fact, many species have genomes that are much larger than our own. The genetic code of mistletoe, for example, is around 94 billion bases long – over thirty times larger than the human genome. Human cells contain two full copies of their DNA, whereas cells of many species contain more. The cells of a brown trout, for instance, contain eight copies.
If we boil down the process of creating a genome to smashing up organic matter to extract the DNA, reading those DNA fragments and putting them back together again in the right order, we start to see why the sheer size of the mistletoe genome poses a challenge – more DNA means more fragments means more complexity.
This simplistic description of course obscures a host of scientific, logistical and economic challenges that have been or are being overcome.
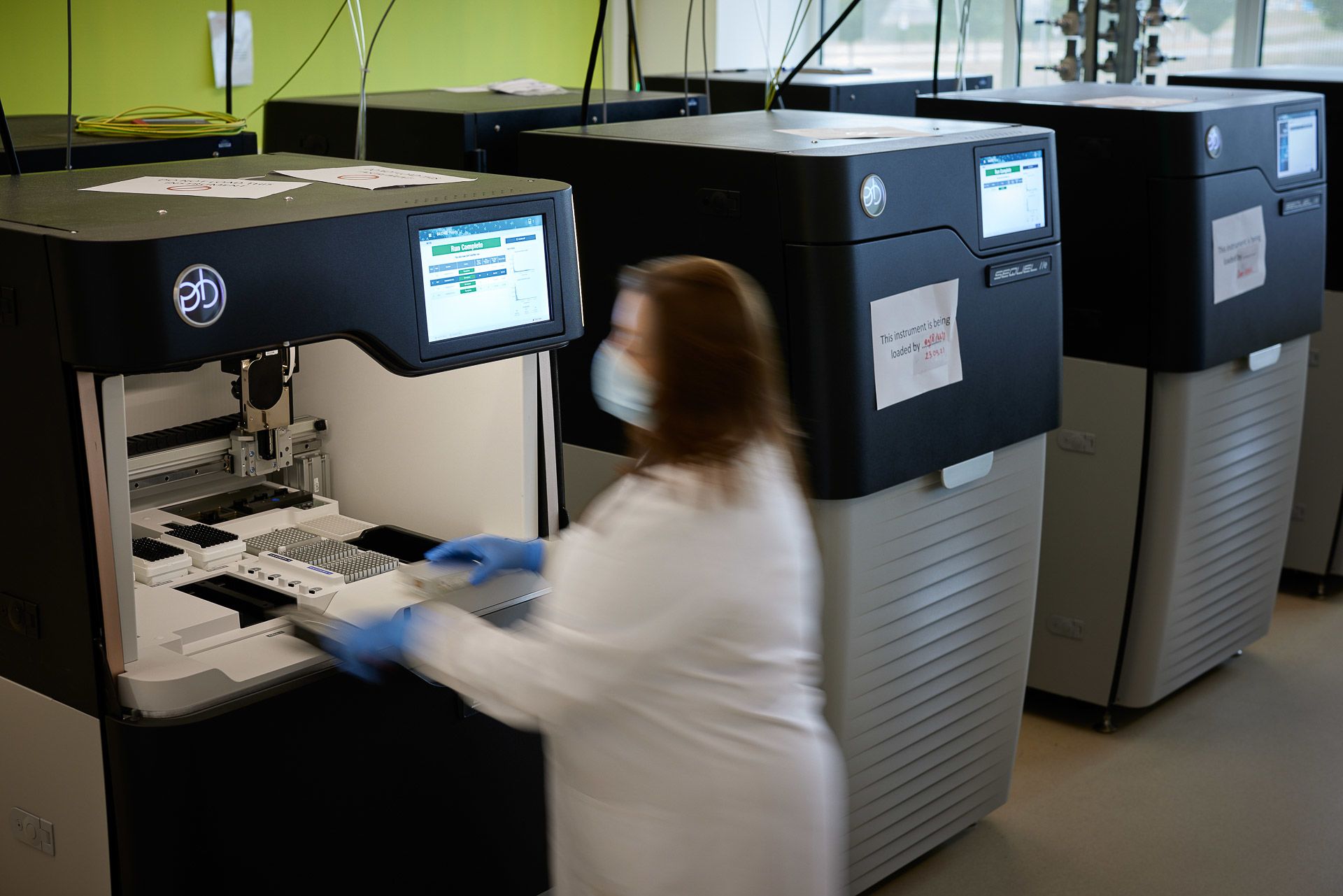
The genome engine
Kerstin’s role today is to oversee Sanger’s novel species sequencing. “Our operation here is very much like an engine. The fuel is the samples that Darwin Tree of Life partners supply. The power output is genomic data. There are lots of moving parts that need to be looked after to keep things running smoothly. But we also have to improve the engine so that we can accelerate towards our goal.”
The first step in the process is to get hold of biological tissue from the target species. Samples are contributed by the ten project partners or one of the many organisations across Britain and Ireland that are helping to collect particular species. All of this activity is coordinated by Nancy Holroyd’s Sample Management team.

“Some partners may only ever send us one sample, usually a very rare species that requires an expert to find and identify. Others, such as the Natural History Museum in London, are sending thousands. With tens of thousands of samples, the process needs to be smooth for all and we need to make sure we are in compliance with wildlife legislation and regulations. There’s a lot to coordinate.”
When samples arrive at Sanger, they’re checked into a sample management system so that they can be tracked through the DNA pipeline, then stored in -70 degree freezers until needed.
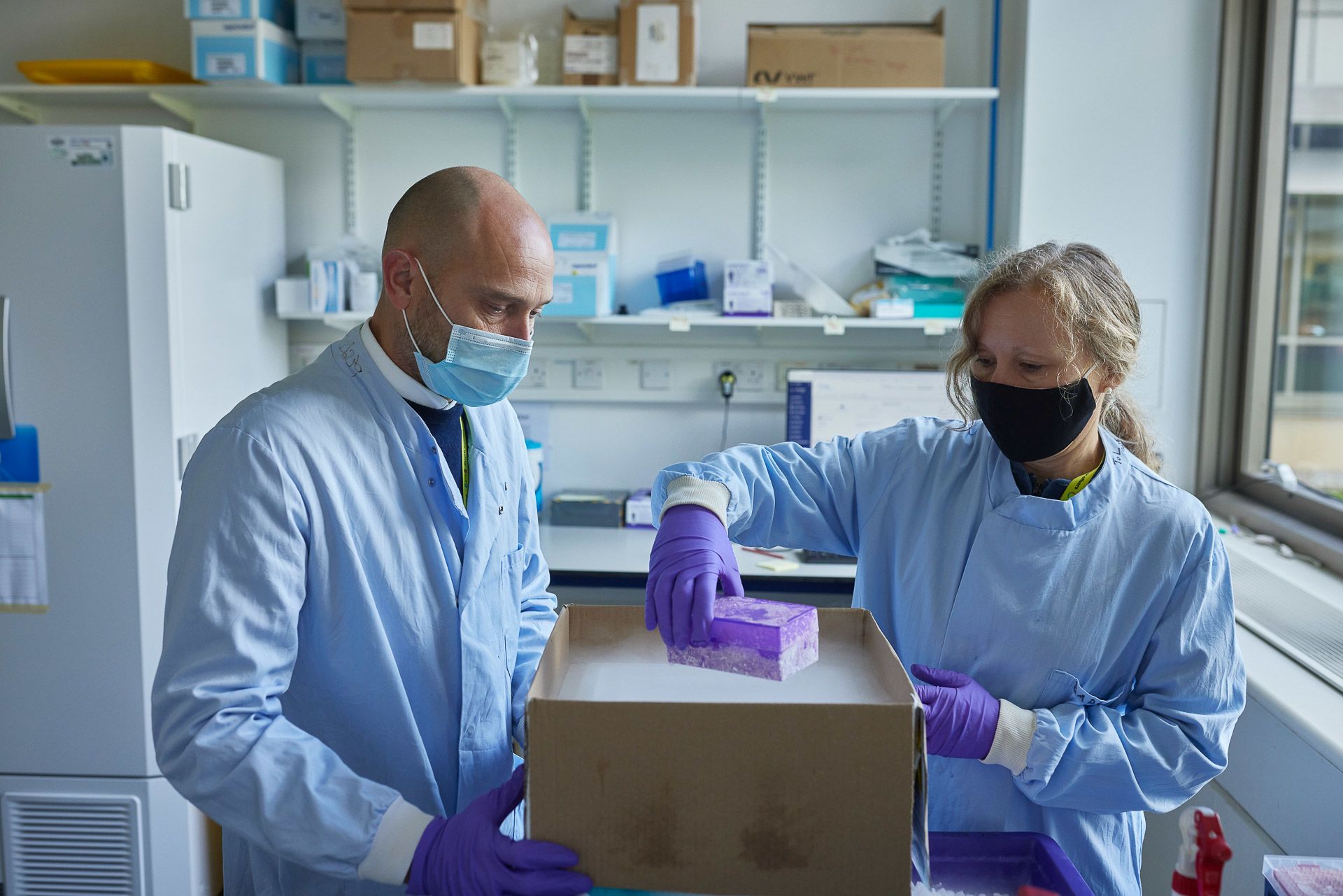
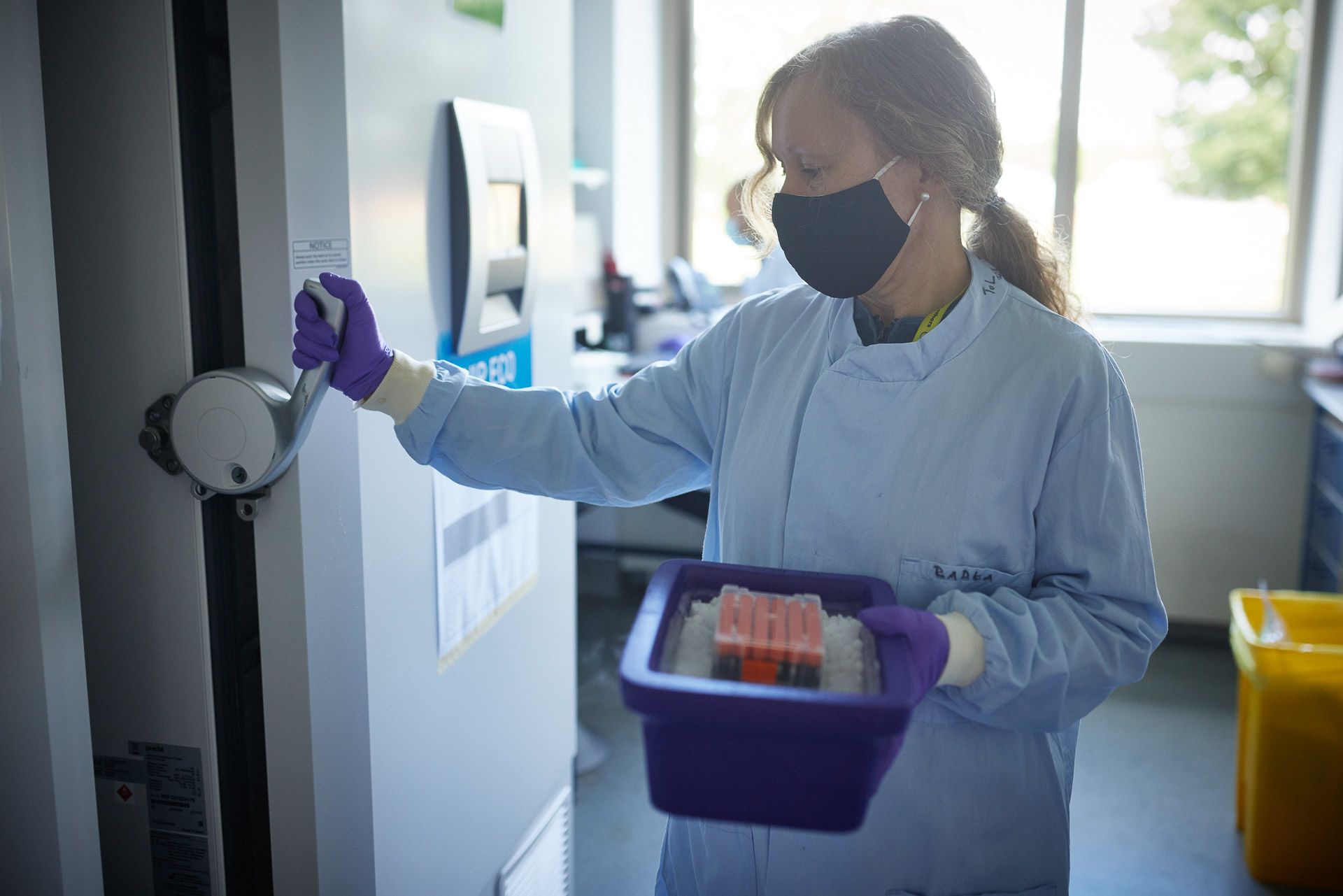




Ian Still and Radka Platte receive Darwin Tree of Life samples
Ian Still and Radka Platte receive Darwin Tree of Life samples


Samples are stored at extremely cold temperatures to preserve the genetic material
Samples are stored at extremely cold temperatures to preserve the genetic material
In the adjacent lab, Caroline Howard’s DNA extraction team is responsible for getting the DNA out of the sample in good condition and in sufficient quantity to provide material for the various sequencing techniques used in the project. Unfortunately, there is no one-size-fits-all process.
“One of the greatest hurdles for my team has been to figure out how to extract DNA in the first place. With many species, such as mammals, we can rely on tried-and-tested techniques. But other species or taxa pose unique challenges. We’ve had to figure out how to get around the slime produced by some molluscs, or the fibrous tissue of some plants.”
The other challenge for Caroline’s team has been to figure out how to extract DNA consistently, to a high standard and in a way that can be scaled up sufficiently in order to create 70,000 genomes in just ten years.
“To sequence this number of genomes we need to standardise scientific processes as much as possible, and we’ve made great progress. When methods do fail, we work together with Sanger’s sequencing R&D team, taxonomic experts and others to find a solution. You test, evaluate and learn from failure. That collaboration has been such a rewarding part of the project, the sense of achievement you get when you solve a problem as a team.”
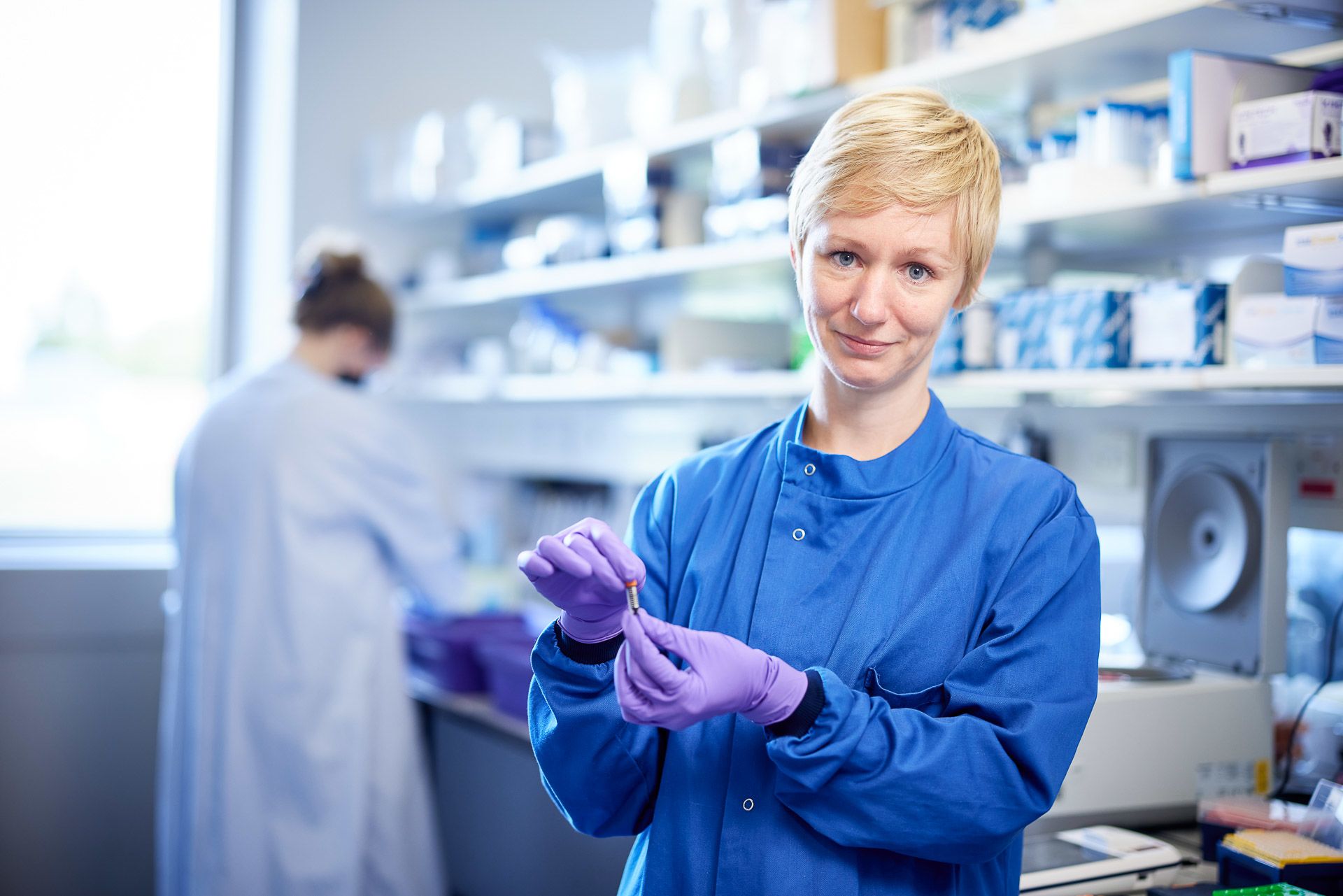
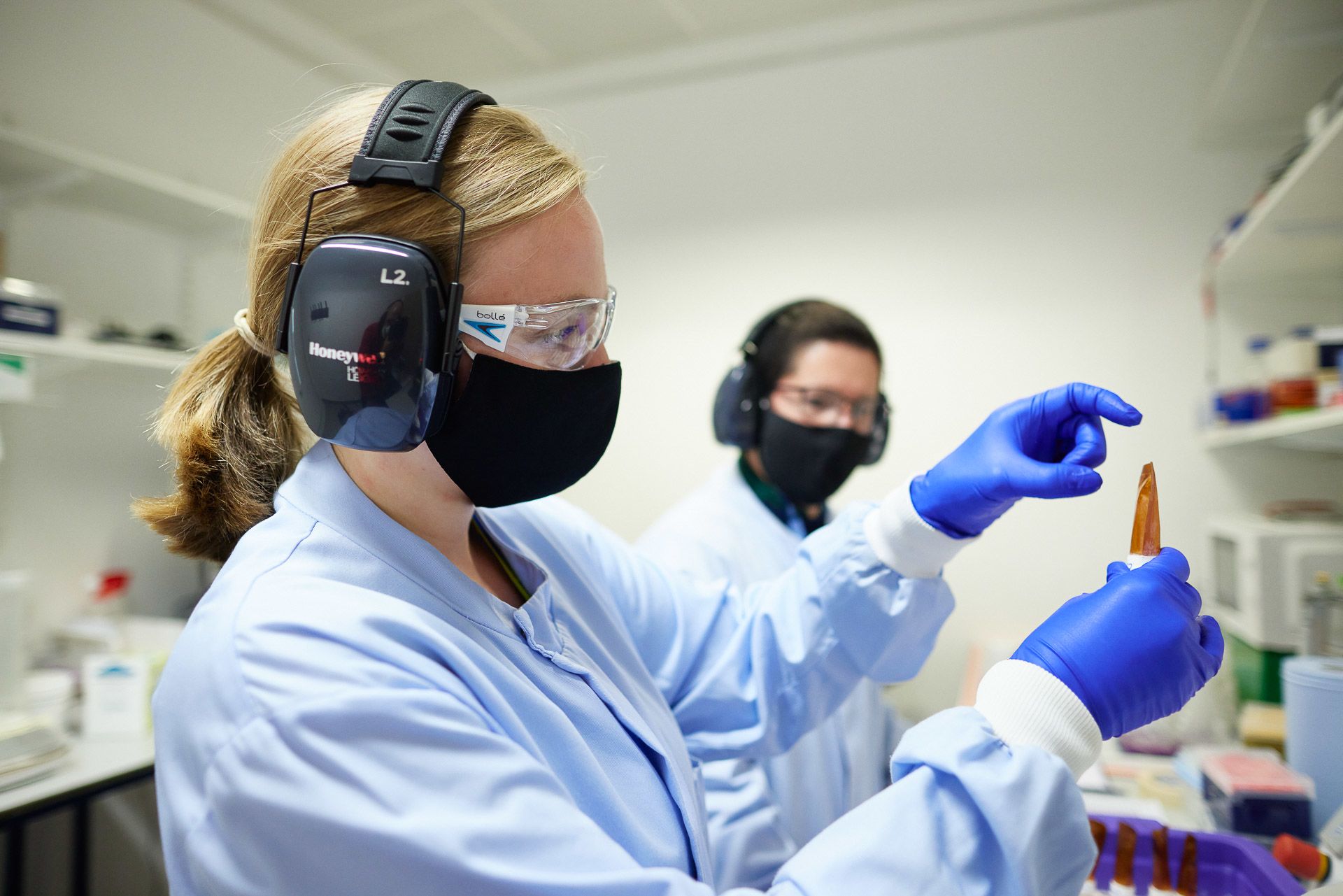
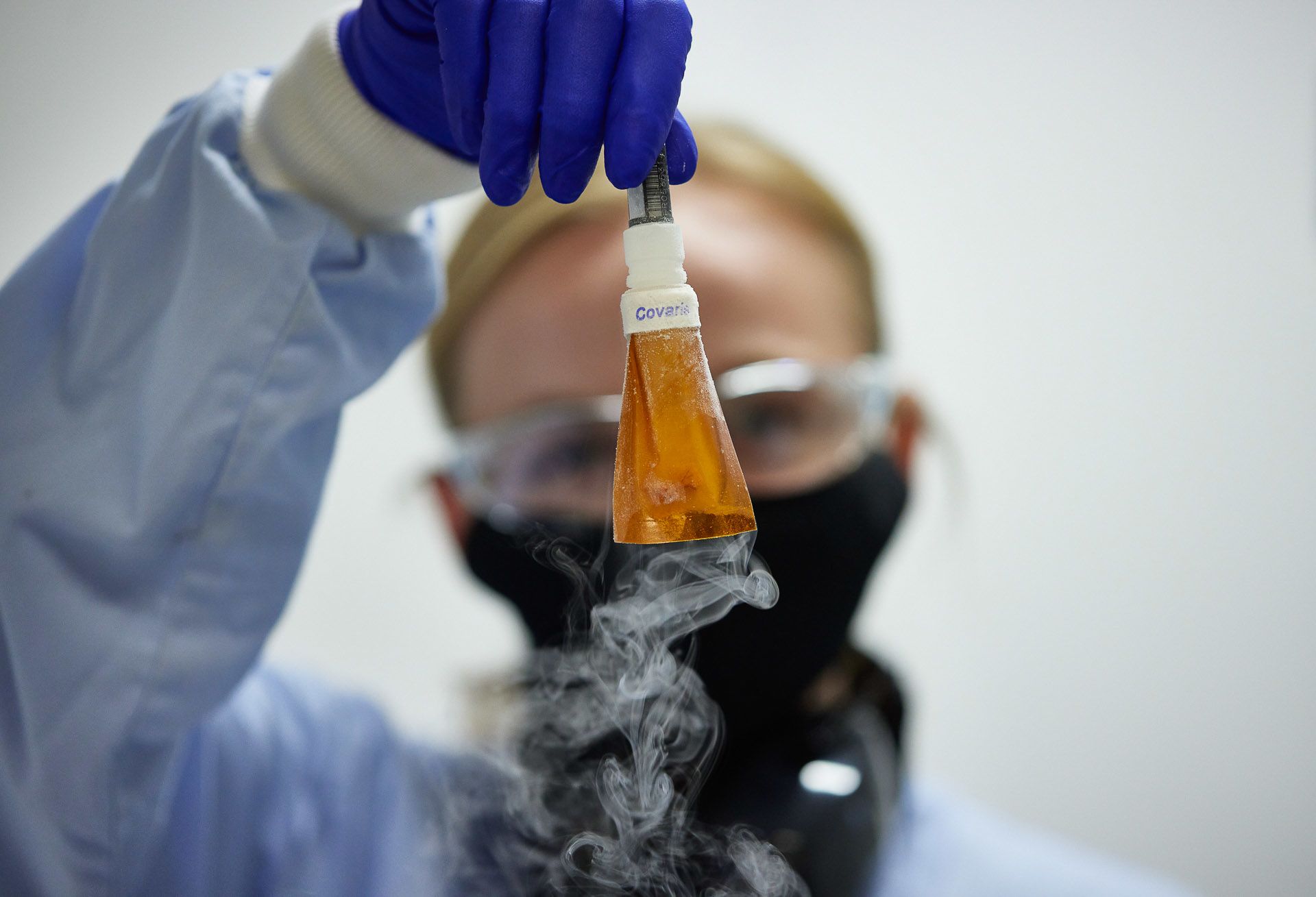
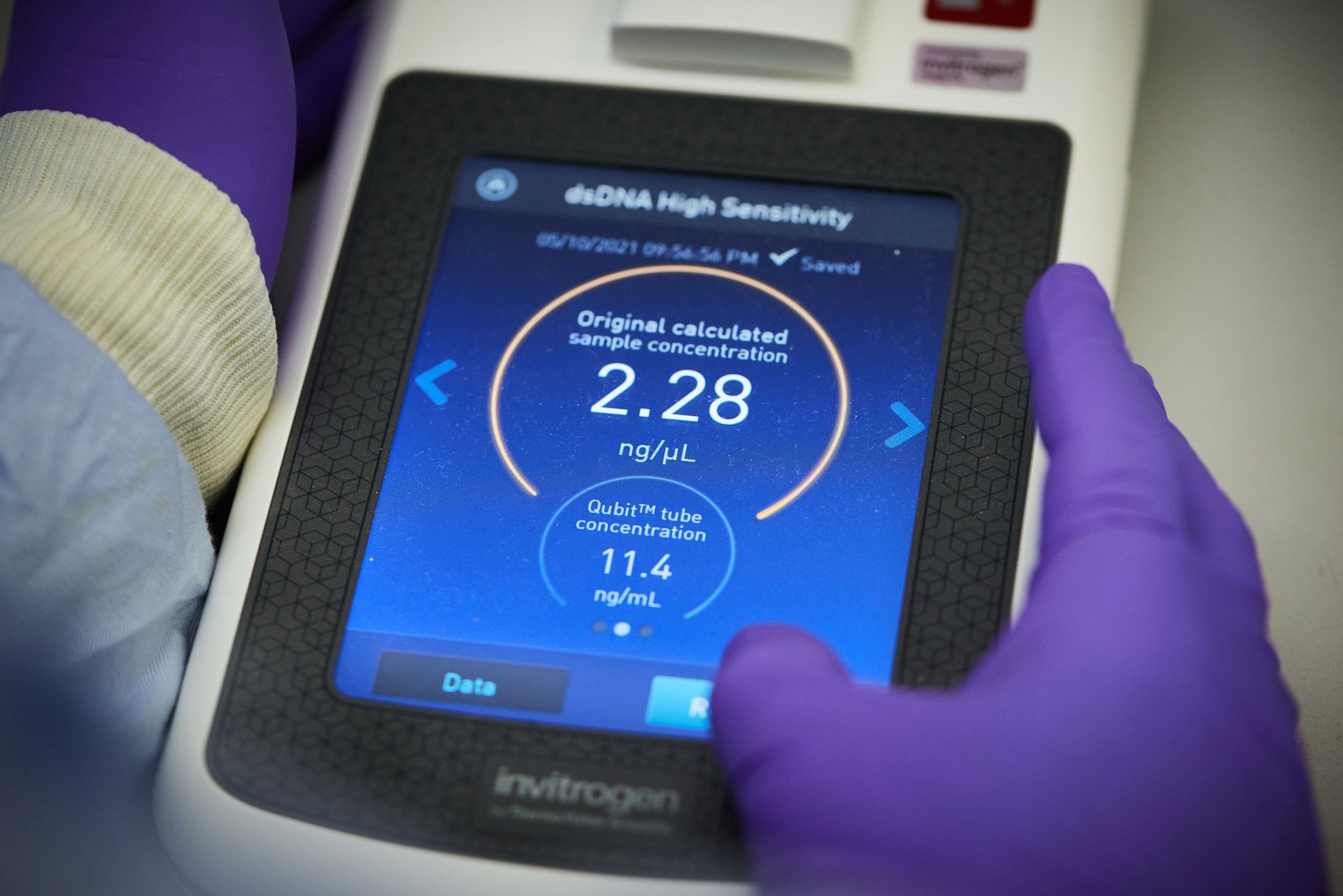






Lucy Kitchin and Juan Pablo Narvaez Gomez flash freeze and pulverise samples, a process known as cryoprep
Lucy Kitchin and Juan Pablo Narvaez Gomez flash freeze and pulverise samples, a process known as cryoprep

Obliterating the sample allows the DNA to be extracted
Obliterating the sample allows the DNA to be extracted

A sufficient amount of DNA is needed for sequencing
A sufficient amount of DNA is needed for sequencing

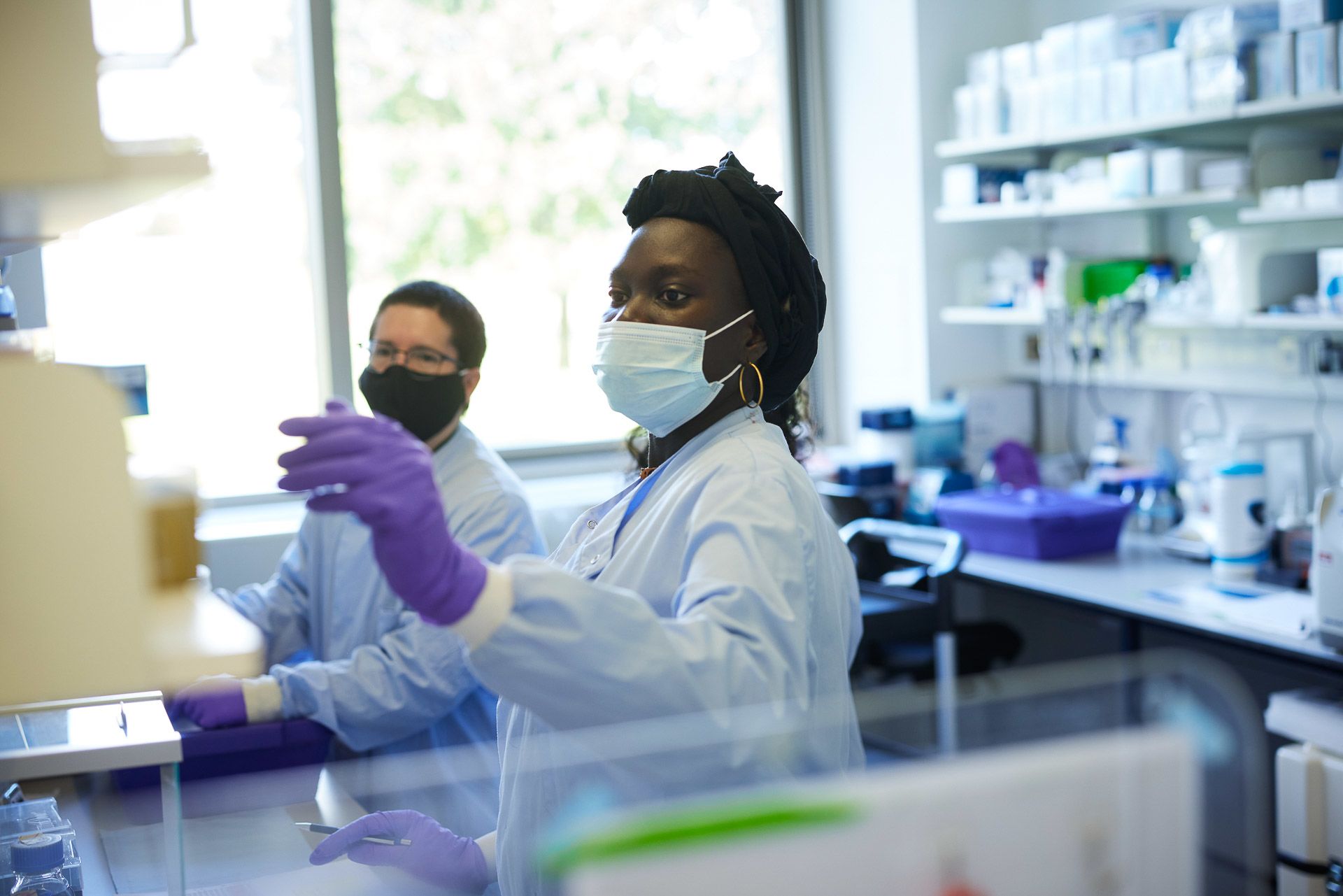
Haddijatou Mbye and Juan Pablo Narvaez Gomez prepare samples for DNA extraction
Haddijatou Mbye and Juan Pablo Narvaez Gomez prepare samples for DNA extraction
Extracted DNA is then sent for sequencing, including to Karen Oliver’s long-read sequencing team. Karen joined Sanger when the Institute was in its infancy, initially working as a finisher on the Human Genome Project, and now works on everything from cancer to conservation.
“It took us over ten years to create the first human genome. Now it’s a matter of days to sequence something new, the speed is just incredible. A big part of my job is knowing what different platforms can do and working with teams across the project to produce the genomic data needed.”
Two kinds of sequencing are used to generate Darwin Tree of Life genome assemblies. The first kind is long-read, where sequences of ten thousand or more DNA bases are generated by PacBio Sequel and Oxford Nanopore PromethION high-throughput sequencing machines. These long reads are the basic building blocks of assembly, like pages in a book. The second kind of sequencing, called chromatin conformation capture or Hi-C, involves gathering long-range information far exceeding the scale of long read data. Using Illumina NovaSeq platforms, Hi-C generates millions of pairs of short reads to indicate links between distant parts of the same chromosome – like footnotes in a book that connect different pages.
“Although we work on many different projects, I think everybody that joins the team is excited about Darwin Tree of Life. It’s the biggest project that we’re working on, a lot of people spend a lot of time thinking about these samples and how to read the DNA within.”
Karen Oliver
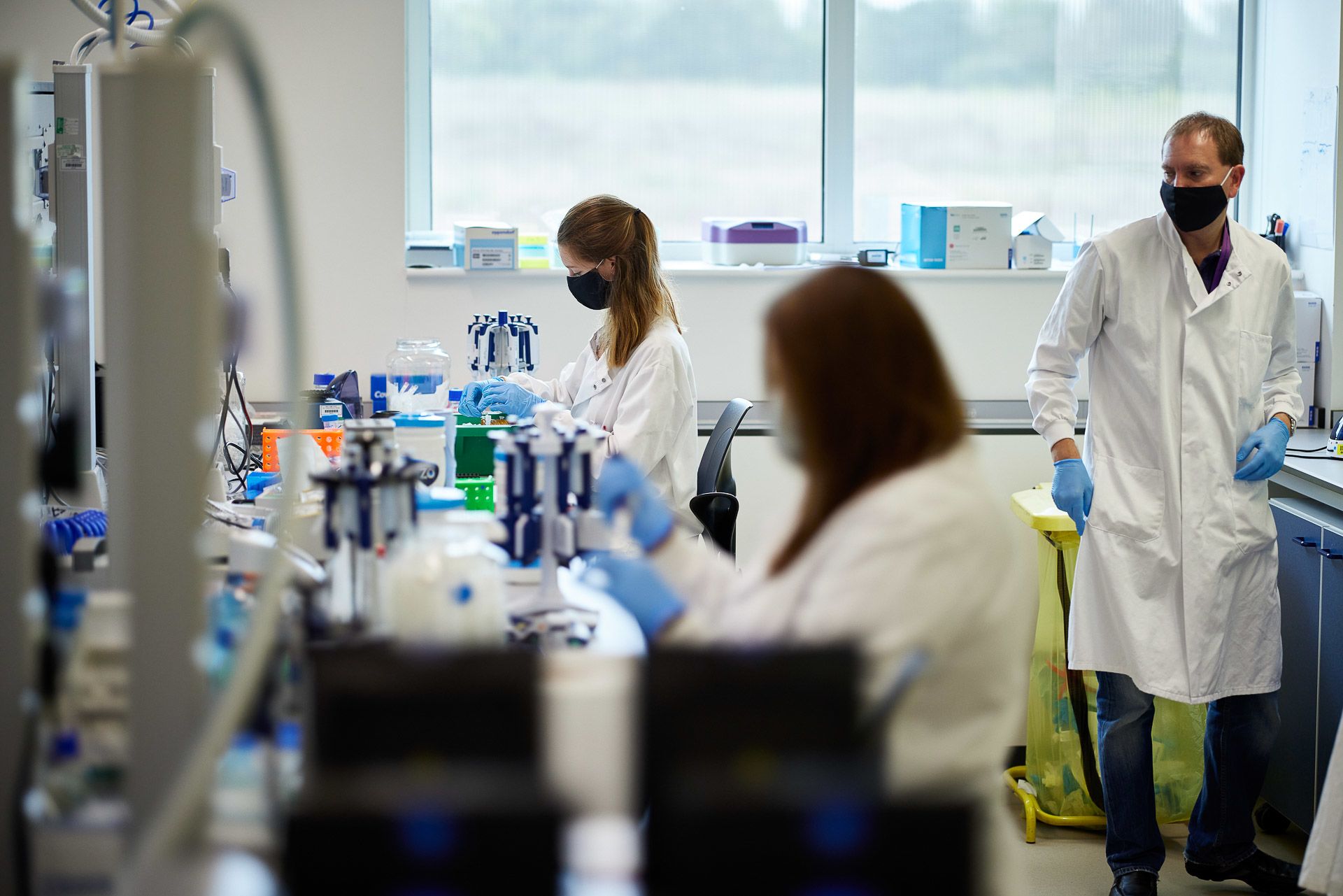
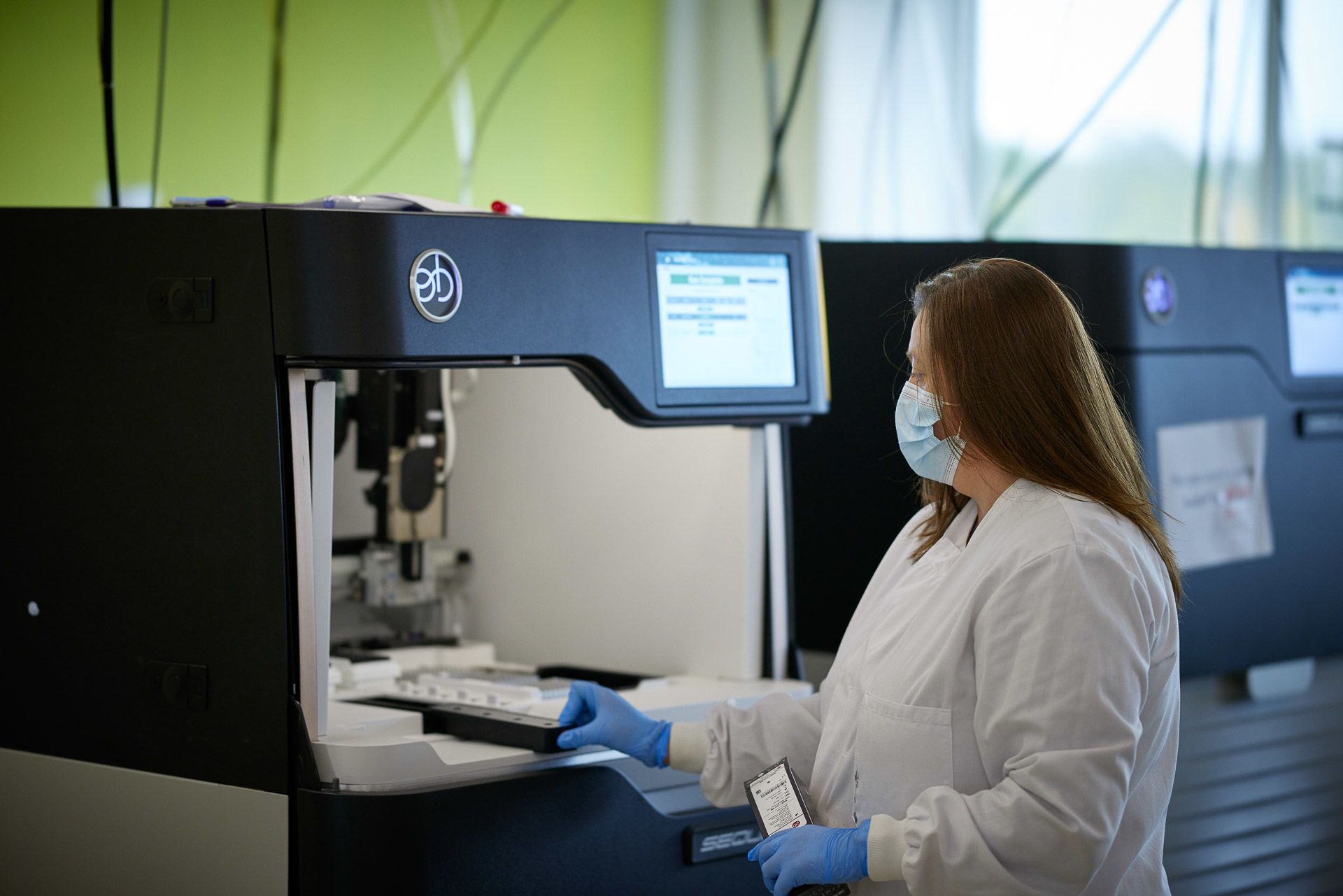
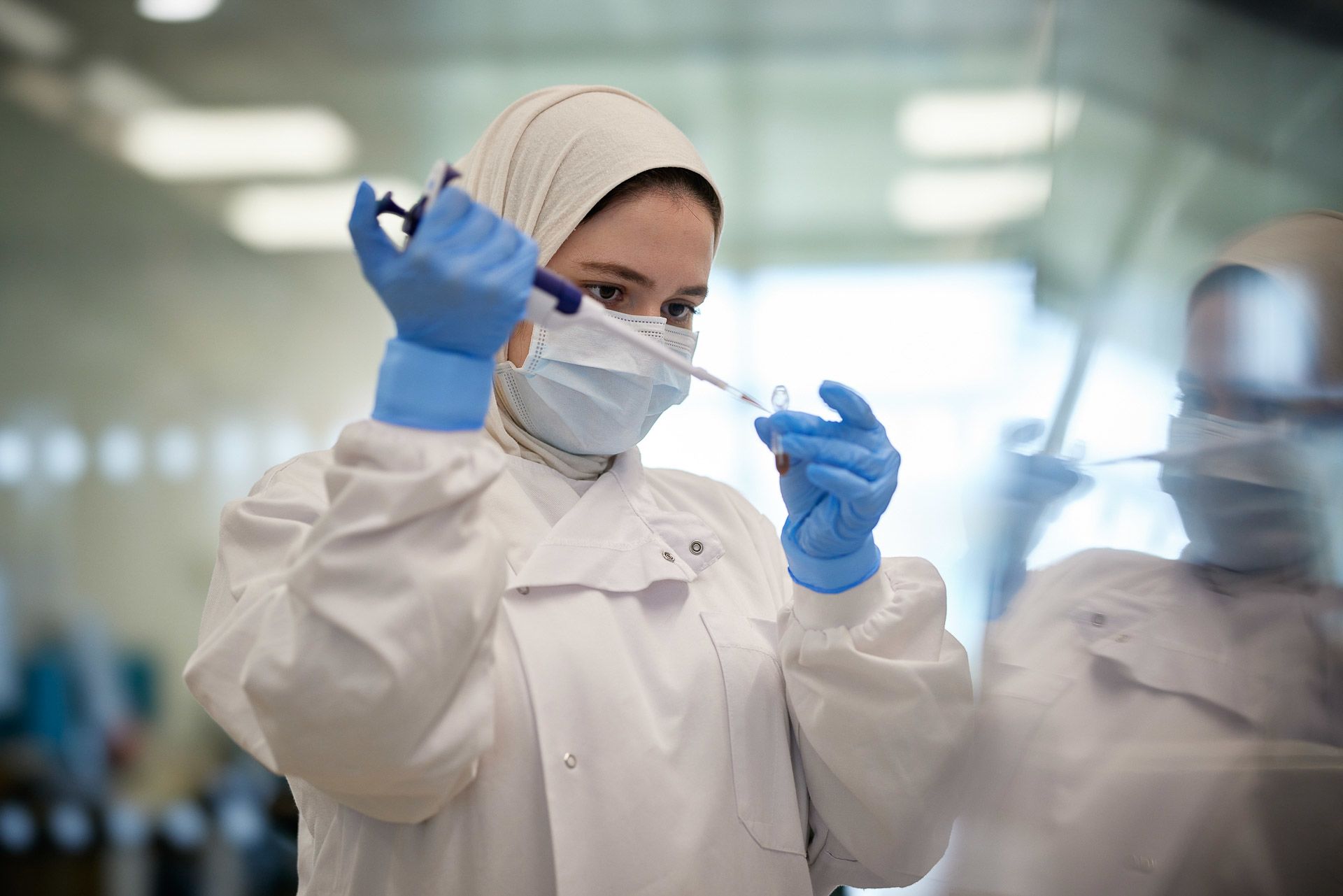





Several manual lab processes need to be performed before samples are ready for sequencing
Several manual lab processes need to be performed before samples are ready for sequencing

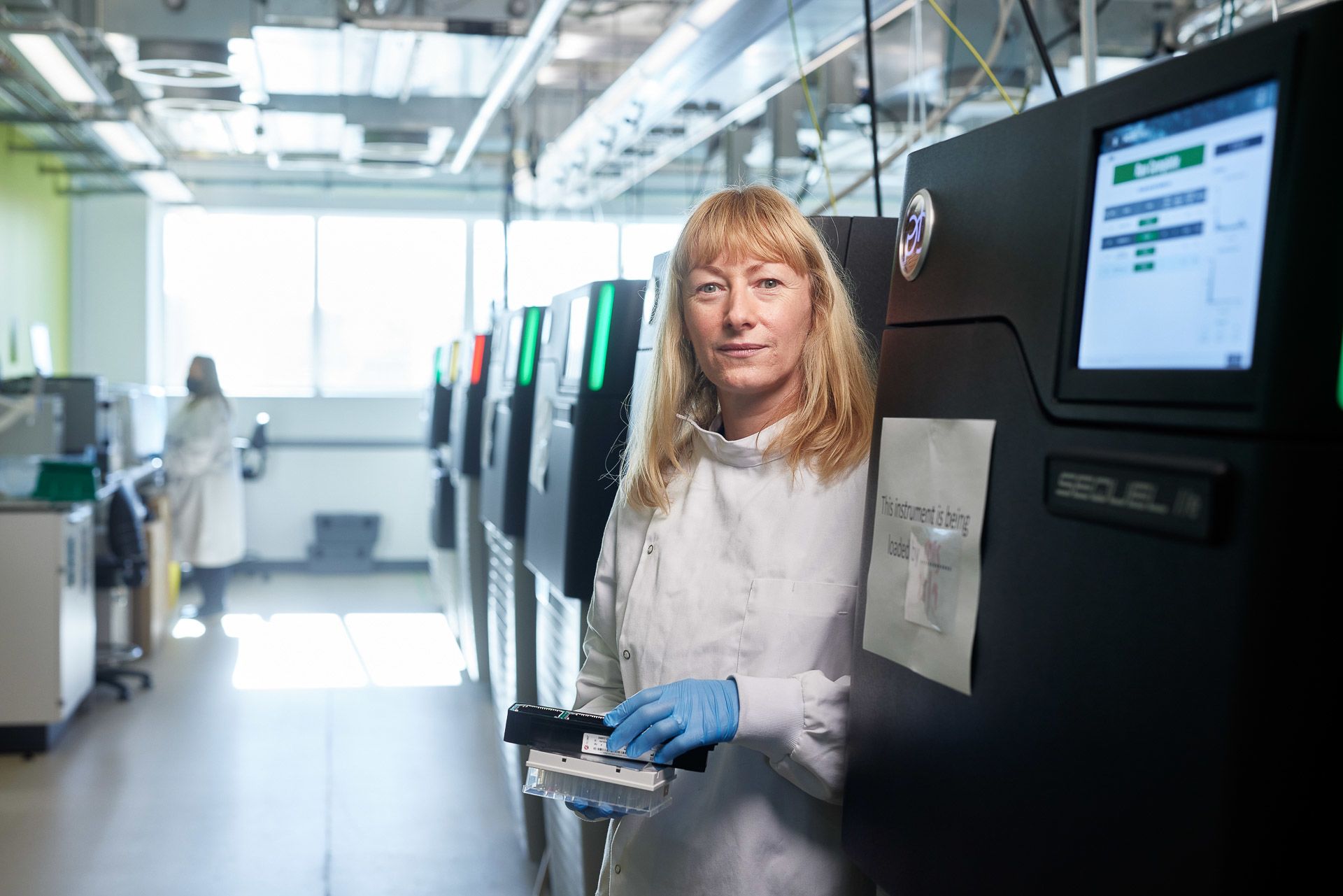
Michelle Smith loads a PacBio long-read sequencer
Michelle Smith loads a PacBio long-read sequencer

Yousra Belattar prepares samples for sequencing
Yousra Belattar prepares samples for sequencing
Picking up the pieces
Reassembling a genome whose DNA has been so thoroughly obliterated is perhaps one of the most impressive feats in the project. Imagine sitting down to do a huge jigsaw puzzle, only to find that much of the picture on the front has been erased. To complicate matters, the mass of pieces belong to multiple copies of the same puzzle. And to top it all off, there may be pieces from a completely different jigsaw mixed in.
The rebuilding of a genome requires several groups of bioinformaticians, the first of which is Shane McCarthy’s assembly team. Shane is a veteran of several ground-breaking genome projects, including the 1000 Genomes Project to map human genetic variation and the Vertebrate Genomes Project.
“One of the first things that we do – and it may sound incredible to be doing this at such a late stage – is to check that we’ve sequenced the right species. Some species are very difficult to identify during collection, so you can have cases of mistaken identity. You can also have genomes within genomes, such as a parasite that was living in the target species when it was sampled.”
Shane’s team create an initial assembly using long-read data, each reporting on thousands of bases of adjacent DNA sequence. “Basically, the longer the contiguous stands of DNA and the fewer fragments that we have, the easier it is to piece them back together. You have more chance of assembling intact genes or chromosomes.” This first round of the assembly process generates a set of DNA sequences that can each be a million or more bases long.
In the second phase of assembly, the long-range Hi-C data are used to arrange the DNA sequence into chromosomes, ordering all the fragments and orienting them. As with other parts of the pipeline, assembly is now semi-automated in order to scale up rapidly in the coming years.
“We can complete a straightforward assembly, such as a butterfly, in a day. We’ve done many butterflies now and have learned lessons that we can apply to other species in that family. But a bush cricket, with a genome that’s three times bigger than our own, may take a week and require more manual intervention. Then you have plants with huge genomes, such as mistletoe, that may demand a different approach.”
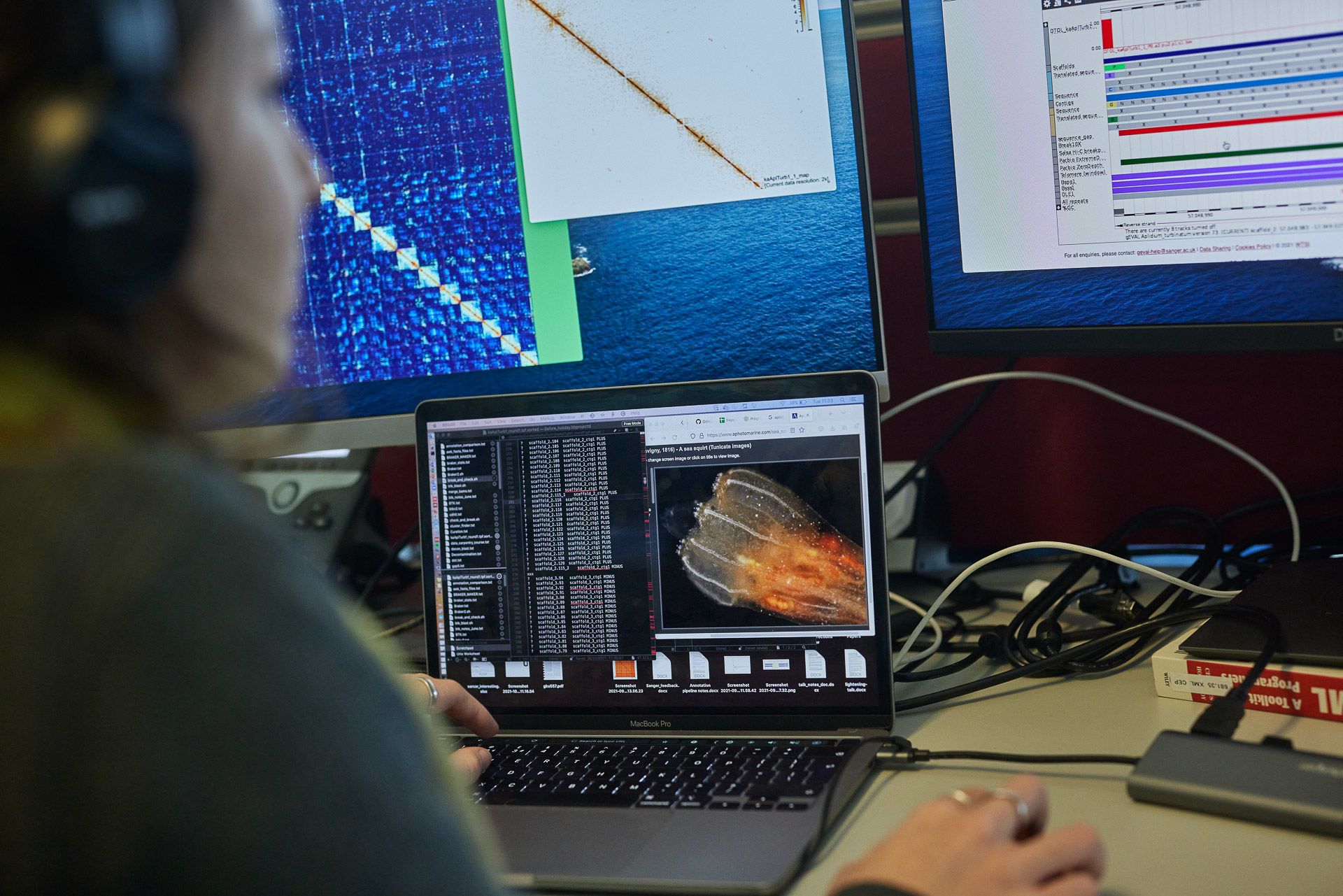
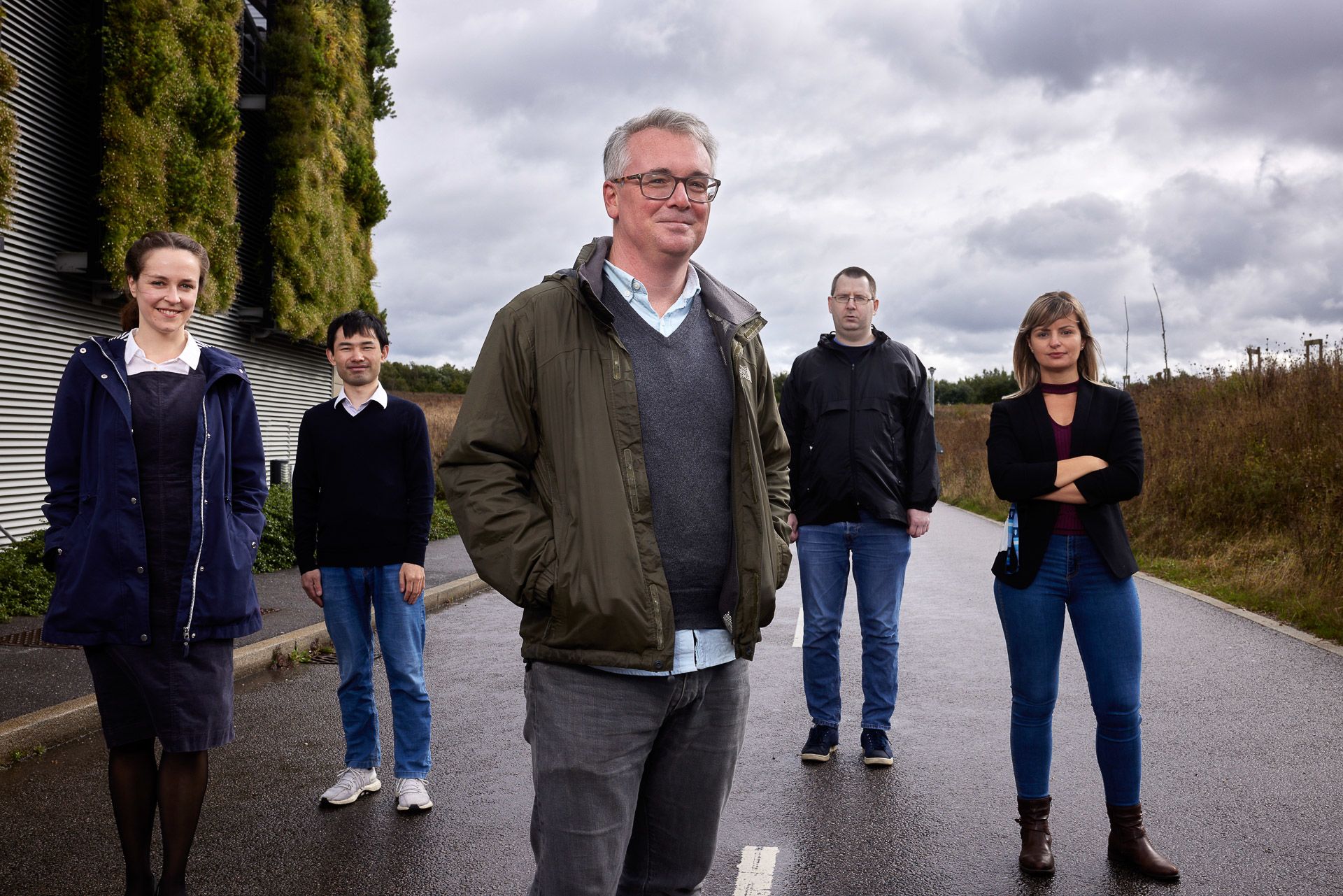


Shane McCarthy and the genome assembly team: (L-R) Ksenia Krasheninnikova, Chenxi Zhou, Shane McCarthy, Eerik Aunin and Marcela Uliano-Silva
Shane McCarthy and the genome assembly team: (L-R) Ksenia Krasheninnikova, Chenxi Zhou, Shane McCarthy, Eerik Aunin and Marcela Uliano-Silva
The issue of genomes within genomes, where the DNA of parasites or bacteria living within the target species become mixed up with its DNA during extraction, is an example of a problem currently tackled by Jo Wood’s curation team.
“Curation is kind of like spell-checking the assemblies built by Shane’s team. We’ll identify errors, such as duplications of a gene or contamination from non-target DNA, then remap the assembly to chromosome level. When the genome leaves us, it is to all intents and purposes whole again.”
Jo is another scientist whose career began working on the Human Genome Project. Back then, genome curation was done manually. Unlike many aspects of leading-edge genomics, it is still done manually today.

“In some ways, my work hasn’t changed since the Human Genome Project. We’re still essentially dealing with four DNA letters in different arrangements. You still have reads, they’ve just got longer. You still have mapping information, it’s just evolved into something new. You still have assembly and scaffolding algorithms – they’ve just got better and can cope with different types of data and more of it. The main difference is purely in terms of scale.”
To give a sense of this scale, analysing the data for a genome with around one billion base pairs of DNA, such as the smoky wainscot moth, requires around one thousand CPU hours and one terabyte of disk space. Manual curation by one of Jo’s team will then take anywhere from two hours to two days, depending on the complexity of the genome.
None of this bioinformatics work would be possible without the vast strides that have been made in computer science in recent years. A lot of work goes on to support the core teams and manage so much raw data, from writing software to developing algorithms to maintaining databases. Much of this work is done by Andrew Varley and Matthieu Muffato, who lead teams that have built bespoke data tracking systems and efficient analysis pipelines to support the whole genome sequencing process, from finding the specimen in the wild to submitting the completed genome to public databases.
Their teams have written more than 250,000 lines of code so far, now deployed on dozens of servers and continuously processing data and metadata. “The challenge,” Andrew says, “is to deal with the inherent complexity of biology. As much as programmers would like to use logic and method to code their way out of problems, they continuously need to adapt to what comes through the genome engine.”
A resource for all
Scale is something that EMBL’s European Bioinformatics Institute (EMBL-EBI) knows all about. The Institute’s mission is to help scientists realise the potential of 'big data' in biology. As one of the ten Darwin Tree of Life partners, EMBL-EBI is responsible for collating all of the data generated by the project and presenting it to the world.
The Genome Analysis team, led by Peter Harrison, develops the Darwin Tree of Life Data Portal, which enables scientists and the public to identify, track and download data generated by the project. The data portal tracks the flow of information, from registering samples to genome annotation, bridging the physical world of the samples to the digital world of the assemblies.
The data are submitted to the European Nucleotide Archive (ENA), one node in the cooperative global system that holds and shares all the public sequence data ever generated. The ENA’s teams, led by Guy Cochrane, receive the data from the Sanger Institute, check it thoroughly, and then integrate it into their database. From there it is shared round the world, and is immediately accessible to all. Submission of the data to the ENA creates a permanent public record of the data, ensuring the project will help drive biodiversity research for decades to come.
To transform the blank text of the genome sequence into a product that includes biological information, another EMBL-EBI team – the Ensembl project – annotates the genome. If we return to the jigsaw puzzle analogy, annotation is a bit like looking at the completed puzzle and beginning to redraw the picture on the front.
Fergal Martin leads the eukaryotic annotation team at EMBL-EBI. “Annotation is really where the genome pipeline ends and analysis begins. We add information to the assemblies, such as the location of genes, which will be useful to the scientists who’ll use the reference genomes in their research.” Annotated genomes, decorated with their genes and repeats, are then uploaded to the Ensembl genome browser, where they can be viewed by anyone in the world.
Storing, analysing and deploying all this information is a big job. The next few years hold several challenges for EMBL-EBI’s teams, not least the volume of data set to be generated. Fergal’s team are also heading into unfamiliar territory in terms of the taxa they are annotating. “Historically we’ve always worked on vertebrates, where there isn’t a huge amount of evolutionary distance between species. Now we are moving on to invertebrates, fungi and plants, where we have less of an understanding about the biological features hidden in the sequence of each genome. You have to learn as you go.”

“The insights these genomes will provide are brand new, nobody has ever sequenced an entire ecosystem on a genomic level before,” said Fergal. “So we’ve never had all the pieces of the puzzle. It is an investment in Britain and Ireland’s biodiversity, and I hope that it gets the recognition it deserves.”
Fergal Martin

A scientific legacy for the next generation

If there is anything that is able to focus attention on the climate and biodiversity crises that we are living through, it should be the thought of the world that we leave to the generations to come.
It is entirely possible that today’s children could be working with Darwin Tree of Life data in 15 years’ time, whether as biologists helping to untangle the evolutionary web that underpins all life on earth, as conservationists helping to preserve this life or as engineers working with new biomaterials. They could be pharmacologists working to unlock the medicinal potential of newly discovered molecules, or racing to overcome drug-resistance in dangerous pathogens.
If they are to have a chance to do so, however, there’s no getting away from the fact that humanity will need to make significant progress on reducing greenhouse gas emissions and preserving biodiversity in the next ten years.
“I have two kids and the climate crisis is definitely a major concern. I haven’t been back to my native Australia for quite a while. My dad goes snorkelling in Western Australia and sees all these great animals on the reefs. I just hope it’s all still there when I manage to go back with my children. As a world we need to do much more on a much bigger scale.”
There are signs that the world is finally getting to grips with these crises. Genomic data will have a role to play in transitioning to net zero, whether that is helping to reduce emissions from food production or replace polluting materials with greener ones.
One thing that is certain – there is a wealth of life and biology waiting to be discovered, as genomics pushes us once again into the unknown.
